Table 2: Equipment List
| \# | Description | Vendor | Price (\$) |
|----|:---|:---|---:|
| 1 | Red laser Light | [Amazon](https://a.co/d/hvr3GUZ) | \$16 |
| 2 | Petri Dishes | | |
| 3 | razor blades | | |
| 4 | Nutrifly (10x 1l) 66-116 | [Genesee](https://www.geneseesci.com/product/nutri-fly-molasses-formulation-pre-mixed-fly-food/) | \$147.67 |
| 5 | all trans-Retinal 25 mg | [Milipore Sigma](https://www.sigmaaldrich.com/US/en/product/sigma/r250) | \$73.2 |
| 6 | fly vials | | |
| 7 | premade fly food | | |
| 8 | fly nap | | |
### Hemilineage
Table 3: Hemilineage
| **Hemilineage** | **Row Labels** | **Sum of c.weight** |
|-----------------|----------------|---------------------|
| 17A | 11404 | 10 |
| 08A | 22549 | 12 |
| | 169711 | 10 |
| 00A | 16388 | 15 |
| | 17047 | 12 |
| 12B | 29239 | 33 |
| 06B | 10812 | 18 |
| | 11682 | 18 |
| 16B | 14424 | 59 |
| (blank) | 10000 | 146 |
| | 10634 | 13 |
| | 24797 | 10 |
| | 225692 | 12 |
| 13B | 11111 | 161 |
| | 35435 | 17 |
| | 156615 | 10 |
| 07B | 14155 | 59 |
| | 16174 | 34 |
| | 16544 | 45 |
| | 18516 | 23 |
| | 20588 | 47 |
| | 101550 | 40 |
| 20A.22A | 11530 | 86 |
| | 11877 | 102 |
| | 13081 | 58 |
| | 16156 | 10 |
| 04B | 15036 | 58 |
| | 16171 | 13 |
| | 16530 | 23 |
| | 16640 | 31 |
| | 16772 | 26 |
| | 19473 | 35 |
| | 21464 | 14 |
| | 22299 | 21 |
| | 101891 | 37 |
| 18B | 13645 | 26 |
| | 13846 | 156 |
| | 14882 | 22 |
| | 17245 | 21 |
| | 17824 | 10 |
| | 18131 | 14 |
| | 21171 | 22 |
| 21A | 15336 | 75 |
| | 16186 | 97 |
| | 16427 | 12 |
| | 17013 | 139 |
| | 17073 | 167 |
| | 17558 | 30 |
| | 18306 | 11 |
| | 20731 | 59 |
| | 20905 | 64 |
| | 24383 | 13 |
| | 27296 | 23 |
| | 28678 | 11 |
| | 36932 | 21 |
| | 101974 | 28 |
| | 152381 | 95 |
| 13A | 14366 | 256 |
| | 15024 | 178 |
| | 16681 | 90 |
| | 17350 | 183 |
| | 27506 | 14 |
| | 33275 | 136 |
| **Grand Total** | | 3291 |
Soffers, J. H., Beck, E., Sytkowski, D. J., Maughan, M. E., Devarakonda,
D., Zhu, Y., … Lacin, H. (2025). A library of lineage-specific driver
lines connects developing neuronal circuits to behavior in the
drosophila ventral nerve cord. *eLife*, *14*.
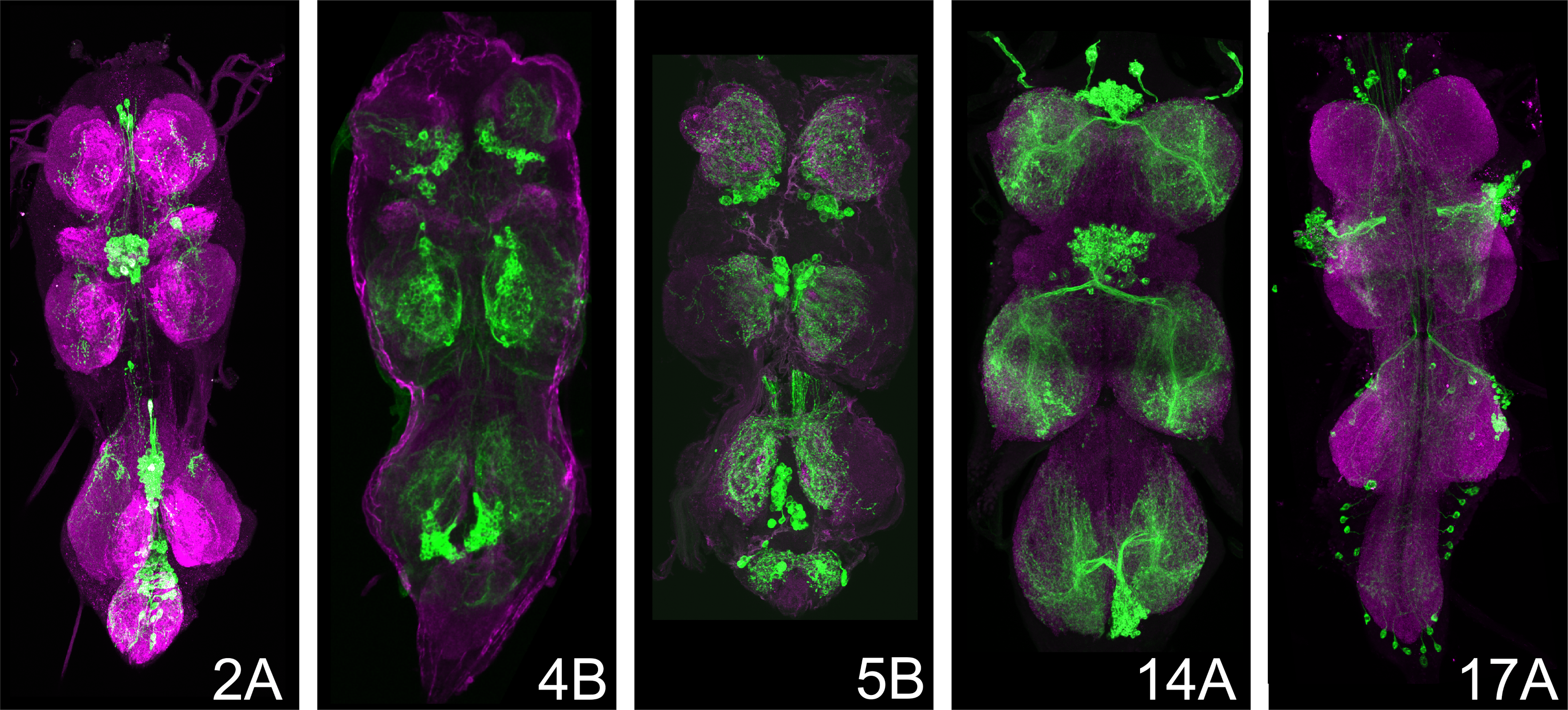 (a)
(a)
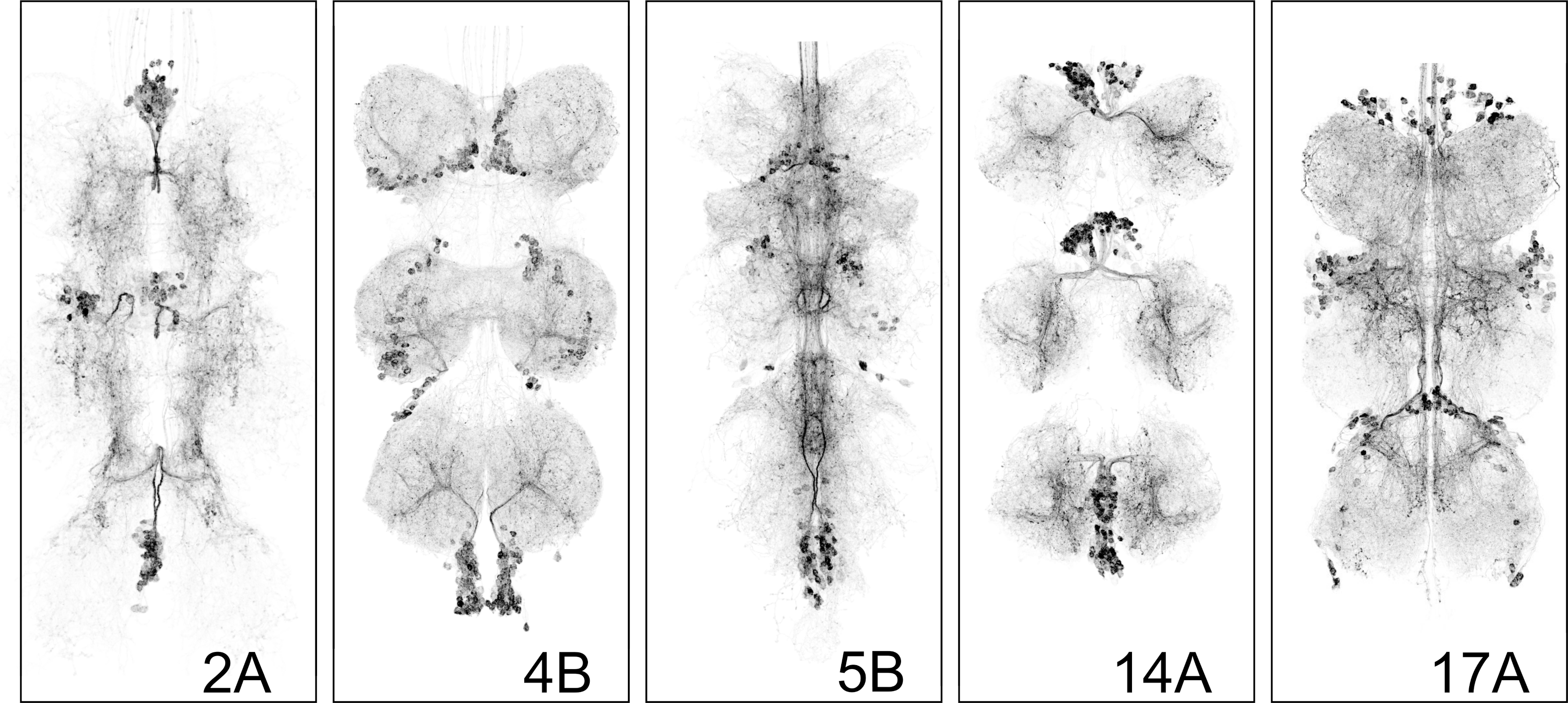 (b)
(b)
 (a)
(a)
 (b)
(b)
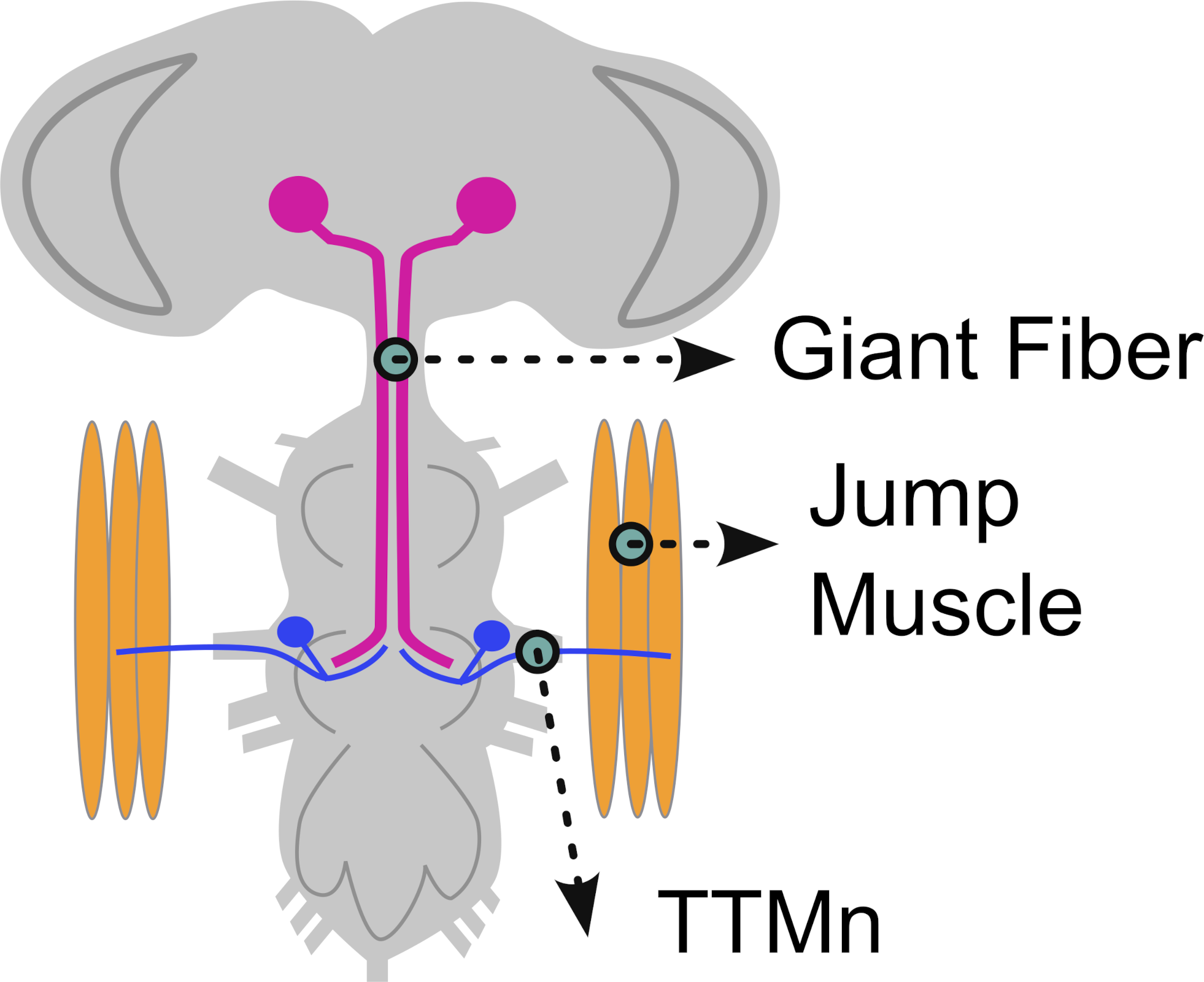 (a)
(a)
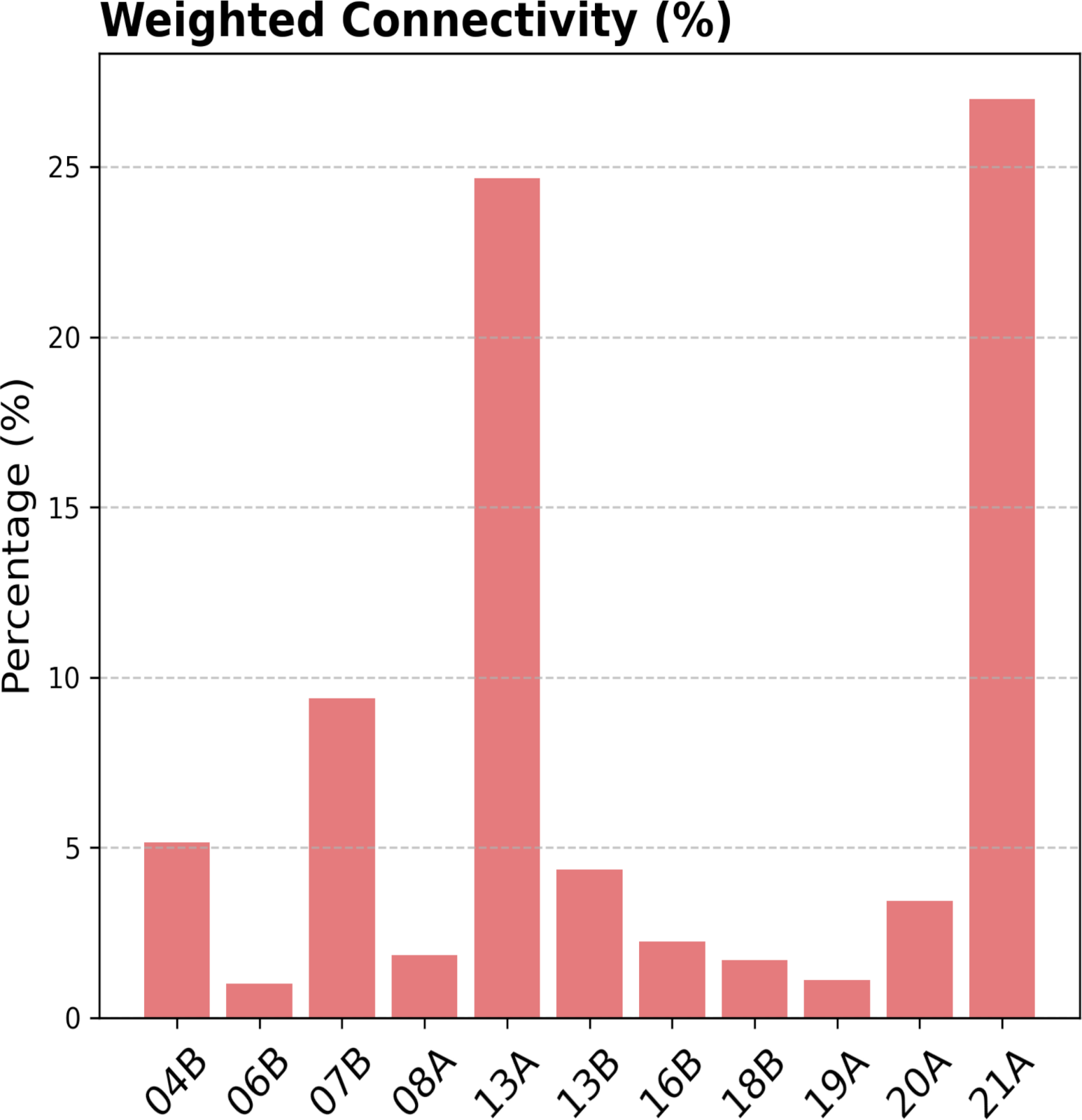 (b)
(b)
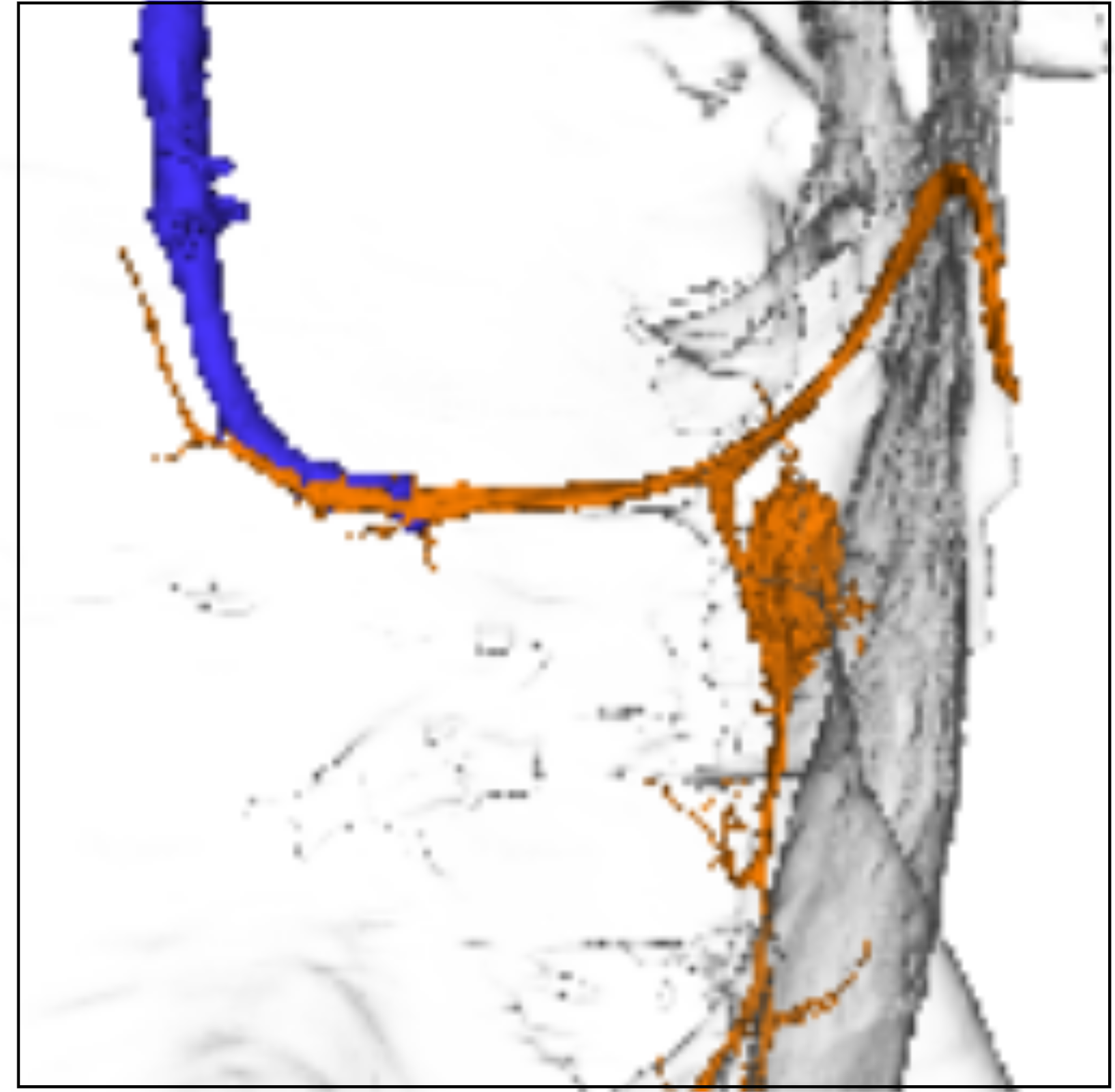 (c)
(c)
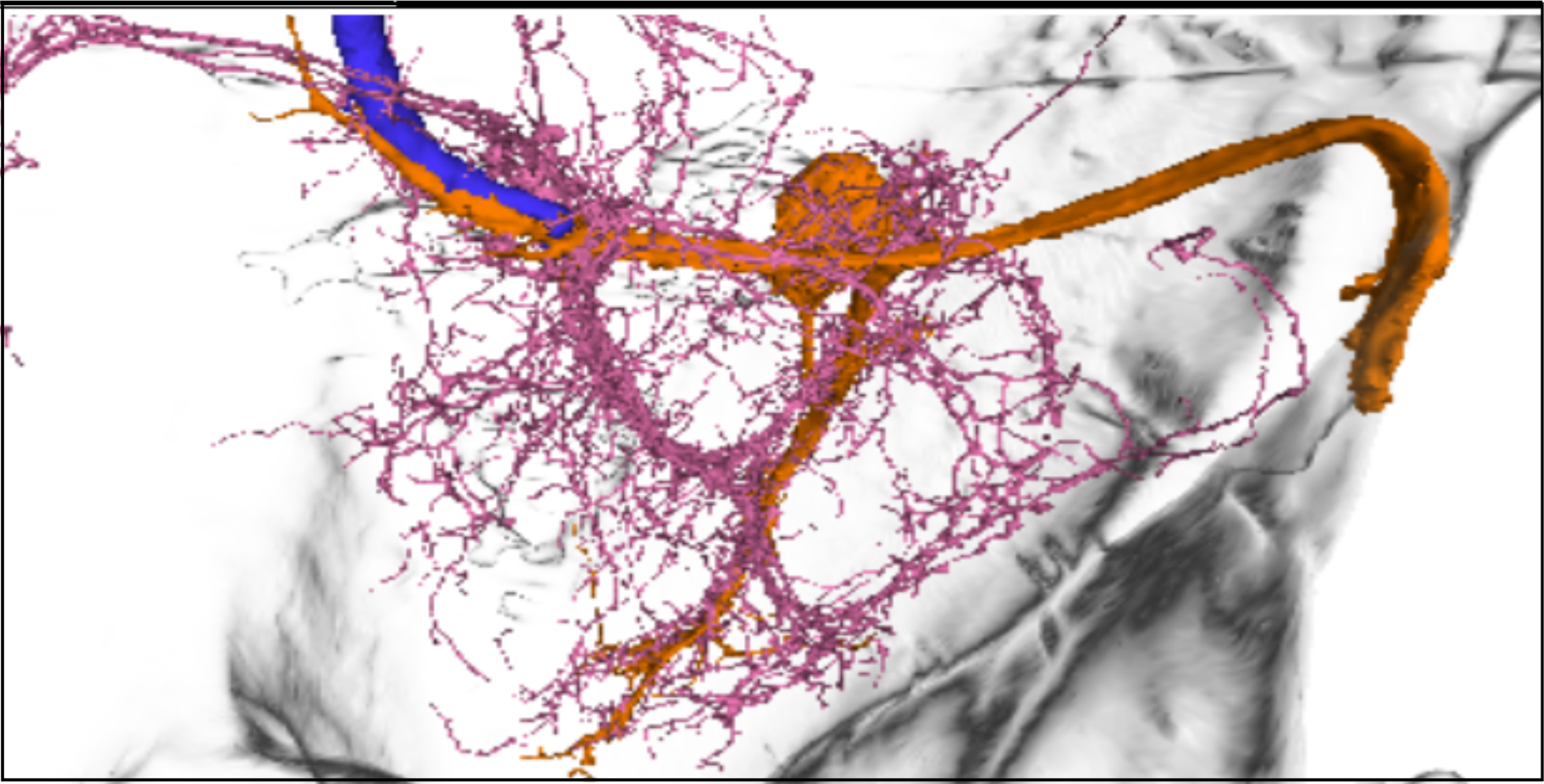 (d)
(d)
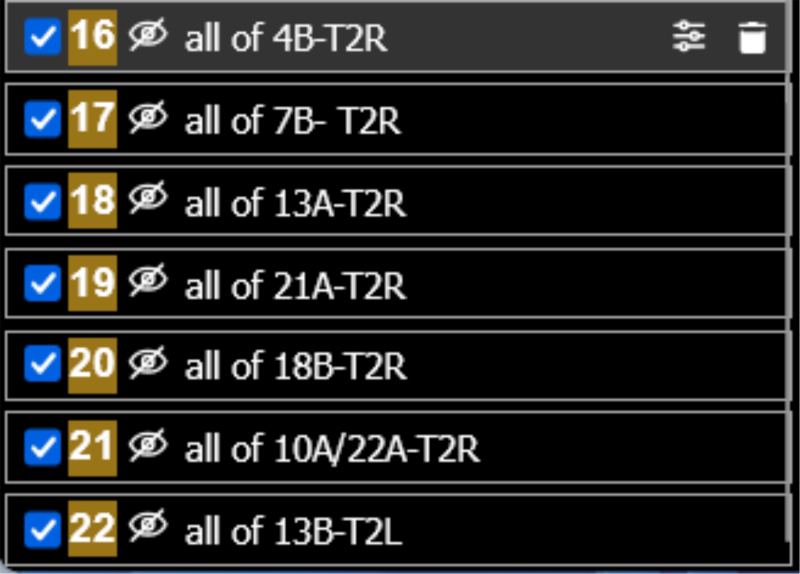
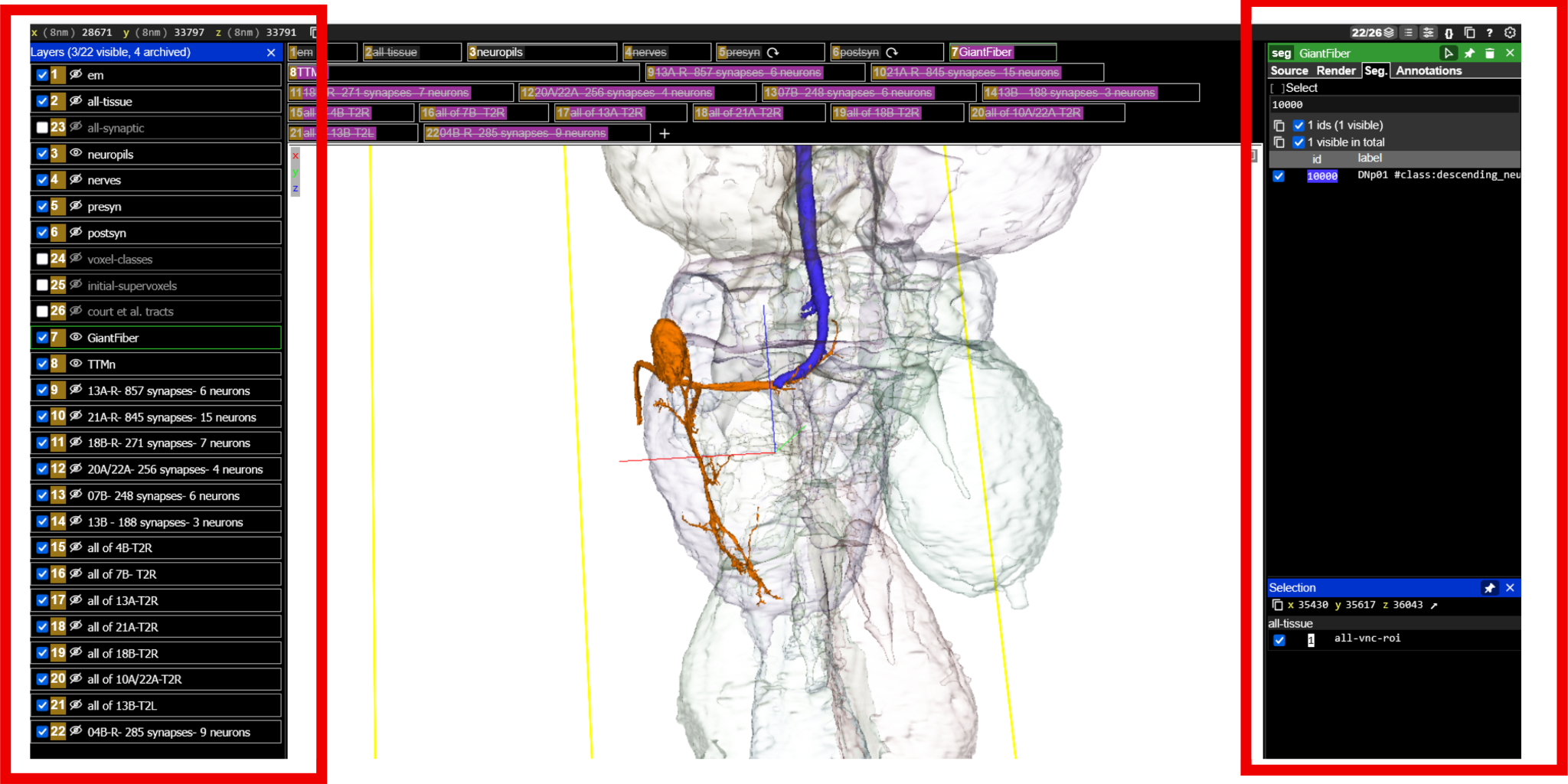 (a) Layer control
(a) Layer control
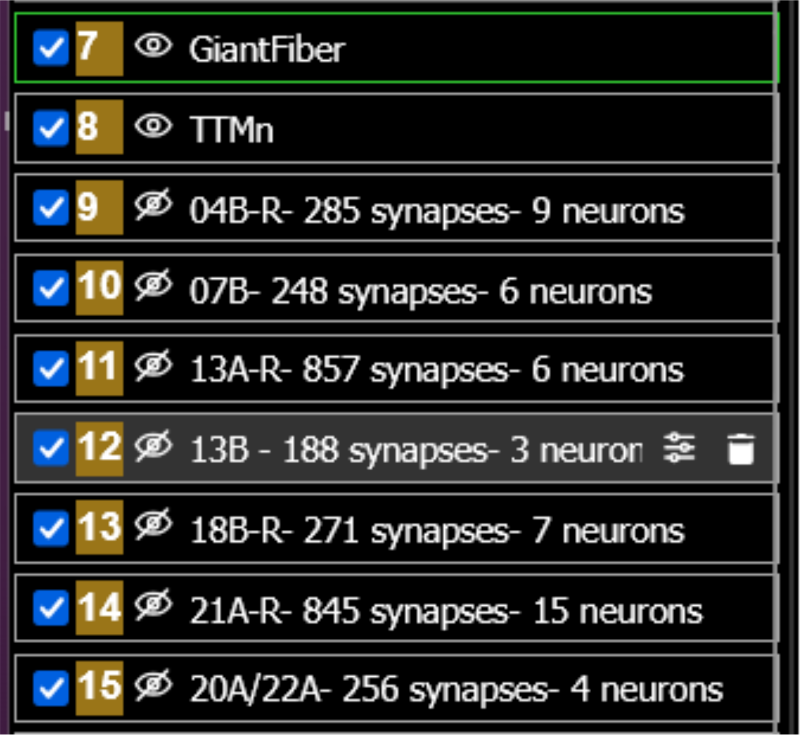 (b)
(b)
 (c)
(c)
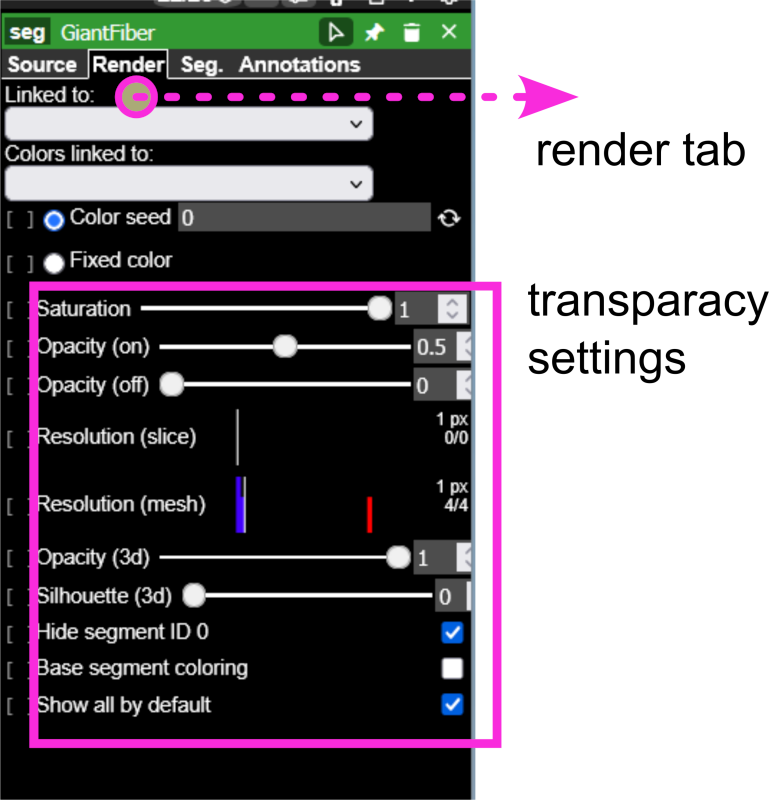 (d) Render tab
(d) Render tab
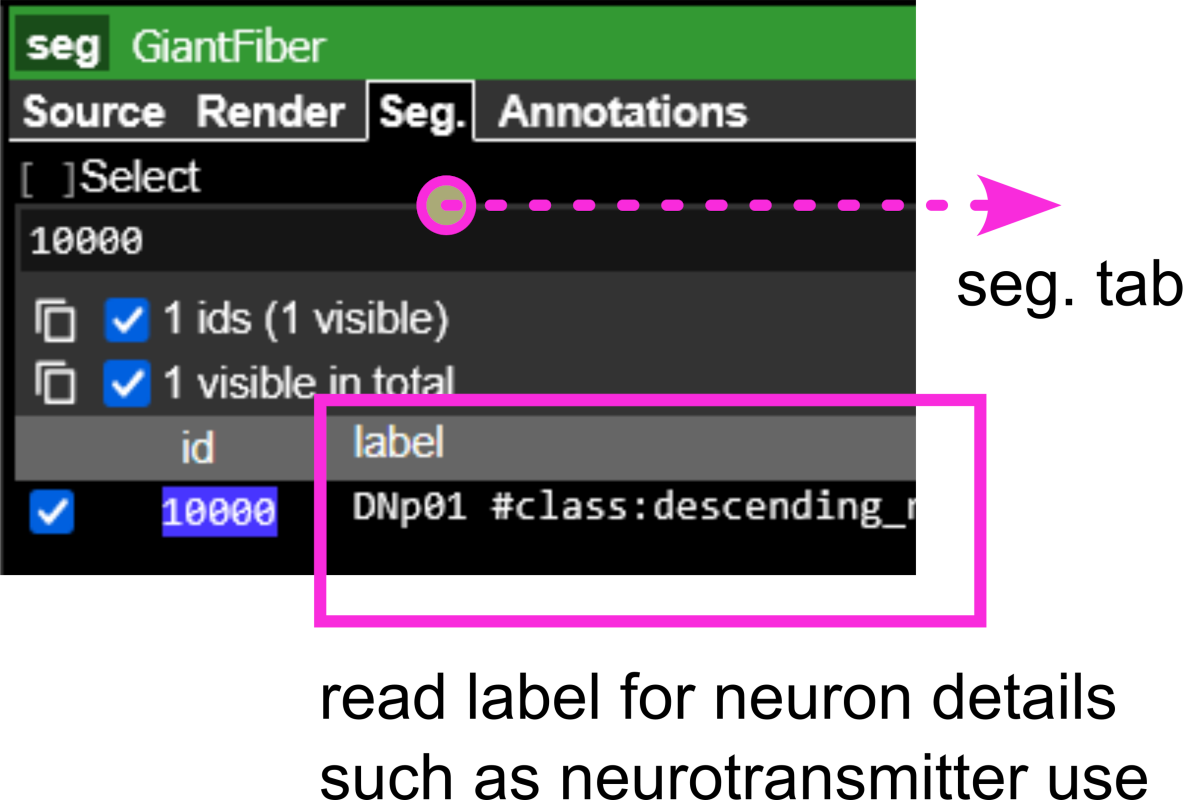 (e) Segmentation tab
(e) Segmentation tab